Researchers from the University of Texas at Austin have developed a method to quickly test hundreds of thousands of potential infection-fighting drugs, which involves constructing bacteria to produce and test molecules that could be self-damaging. This research comes at a pivotal time, for humans are increasingly becoming immune to existing antibiotics.
The method is described in the journal Cell.
“Our arsenal of antibiotics is quickly dwindling and there aren’t many ideas out there to find new ones,” said Ashley Tucker, a postdoctoral fellow at UT and leader of the experimental work.
“Most of the low hanging fruit with regards to natural products have already been picked and now companies are turning to derivatizing old classes of antibiotics that have been around for the last several decades,” she continued. “We wanted to come up with a better way to quickly screen many different chemistries of antibiotics in order to find new scaffolds with antimicrobial potential.”
The World Health Organization claims that antibiotics have increased the average human lifespan by about 20 years, but now the drugs’ effectiveness is dwindling. The U.S. Center for Disease Control has found that at least 2 million people in the country suffer from antibiotic-resistant infections every year.
In an effort to find new types of infection-fighting drugs, lead researcher and assistant professor of molecular biosciences Bryan Davies, and his team, screened close to 800,000 molecules, or peptides, to see if they could kill harmful bacteria.
The researchers found that several thousand peptides killed E. coli bacteria, and that one of those molecules, named P7, can kill other forms of pathogenic bacteria and is safe in mice.
This new method called SLAY (Surface Localized Antimicrobial Display) enables researchers to use less time and money when screening hundreds of thousands of peptides. They hope that it will eventually be used as the standard tool for finding more antibiotics.
“SLAY is a method that puts the bacteria to work for us,” said Tucker.
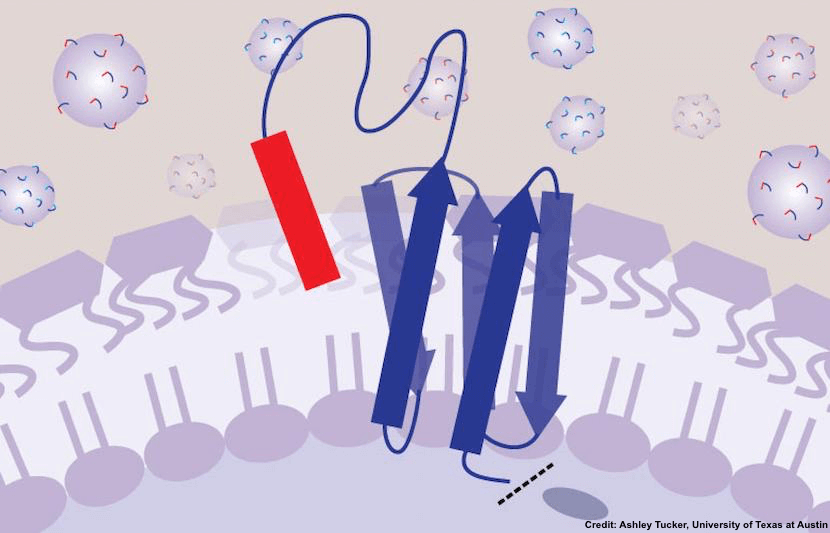
SLAY allows the bacteria to make their own genetically encoded peptide and to tether the peptide to its surface, similar to how a tetherball is attached to a pole. One end remains fixed to a cell membrane, and the other end remains free to float around and touch the bacterial cell surface.
“If the tethered peptide kills the bacteria, we are able to figure out the genetically encoded peptide sequence using next-generation sequencing,” said Tucker. “We then synthesize the individual peptide and perform minimal inhibitory assays on a panel of bacteria to confirm its antimicrobial properties.”
This new method could provide a huge advance in efficiency compared to previous slower and more expensive methods.
“Current screening methods limit us to testing only a few thousand compounds at a time,” said Tucker.
“SLAY gives us, not just the higher throughput, but also a biologically relevant means to screen extensive libraries of peptides for development. We were able to screen approximately 800,000 peptides in one 5 mL test tube faster, cheaper, and more effectively than any current drug-screening platform on the market today.”
Because the researchers found that P7 is capable of killing pathogens, they plan to create thousands of derivatives, or small variations of the molecule, and run them through the SLAY screening test.
Additionally, the team is working closely with Avalon Ventures, a company based in San Diego, California with a long-standing and successful focus on seed and early stage companies, to transform SLAY into a company, said Tucker.
“We plan to expand the technology as well as to continue developing and optimizing current leads,” she said. “It is fairly trivial to find a peptide that kills bacteria. It’s much more difficult to find a peptide that won’t kill a mouse. We are moving forward into animal testing, which, so far, has proved promising.”